Middle-level Analysis of a BiTE Molecule
Efficient digestion of multi-domain BiTE molecules, separating the domains for high-quality LC-MS analysis and precise localization of protein modifications

Bispecific T-cell Engager (BiTE) molecules are a type of fusion protein therapeutics where two scFvs are fused, forming a bispecifc molecule with one scFv targeting cytotoxic T-cells, and the other one targeting a cancer antigen. The protein consists of four protein domains, the two scFvs both being made up of a VL and a VH region, joined by a total of three flexible GS linkers. The large size of this protein, 54 kDa, may pose a challenge for product quality monitoring by intact mass spectrometry since standard MS instrumentation may not be able to provide monoisotopic resolution of the protein.
A research grade biosimilar of blinatumomab, a BiTE molecule with specificity towards CD19 and the T-cell marker CD3, was digested with GlySERIAS Immobilized overnight at room temperature. Direct analysis of the digested sample by LC-MS showed that all three linker regions were digested, with the α-CD3 VL and VH chains eluting as one chromatographic peak, and the α-CD19 VH and VL chains eluting as separate peaks. A small peak from the linked α-CD3 VH and α-CD19 VH domains revealed that this short linker was not completely digested, indicating that GlySERIAS has a lower activity on short linkers. The separation of the four domains provided high quality spectra with monoisotopic resolution, as compared to the intact protein analysis. A few different variants of each domain could be observed, with different numbers of linker residues remaining.
Efficient linker hydrolysis in the BiTE molecule
The α-CD3 VH domain could not be fully baseline resolved from the α-CD19 VH domain during chromatographic separation, resulting in some co-elution which could be observed in the associated mass spectra. The separation of the four domains yielded information about the location of protein modifications identified on the intact protein level. For example, a protein modification of 37 Da, which was also identified on the intact level, was assigned to the α-CD3 VH chain. Furthermore, two modifications of 240 Da and 320 Da, respectively, were assigned to the α-CD3 VLdomain. The latter also contained a His-tag with five or six histidine residues. These data demonstrate the power of middle-level workflows using GlySERIAS Immobilized for quality control of fusion proteins containing several flexible linkers.
Middle-level analysis improved mass spectral resolution
Protein modifications can be localized at the subunit level
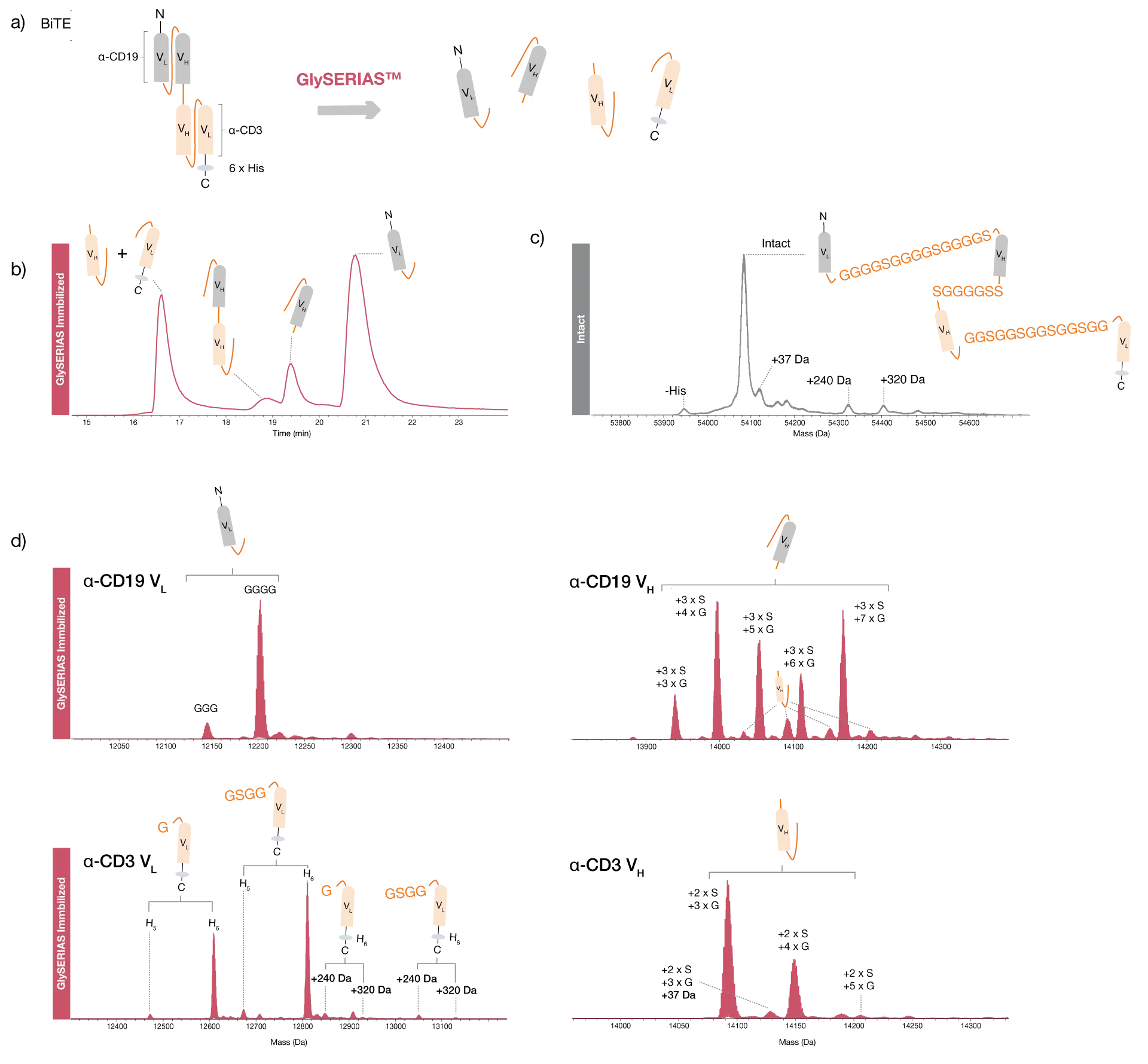
Related Products
Resources
GlySERIAS®
Download Product Folder