Enhanced CE-MS analysis of bispecific and LALA antibodies using SmartEnzymes
Gstöttner et al. used SmartEnzymes for their CE-MS analysis of highly homologous bispecifics, one of them containing LALA mutations.
Bispecific antibodies (BsAb) offer an attractive alternative to traditional mAbs since they can bind to two different epitopes simultaneously. Because they are composed of four different peptide chains, rather than two, they are subject to additional macro- and microheterogeneity which requires proper analysis and monitoring. Bottom-up approaches together with mass spectrometry (MS) have been employed to address such heterogeneity but such methods are unable to handle minor mass shifts and does not provide domain localization information.
In a new article by Gstöttner and colleagues at Leiden University Medical Center together with Roche Penzberg, sheathless capillary electrophoresis (CE) coupled to MS was used to probe two highly homologous BsAb. They looked at both intact BsAbs and their subunits generated using SmartEnzymes. FabRICATOR was used for below hinge digestion in one of the BsAb while the second BsAb proved to be more challenging due to a LALA mutations and required above hinge digestion using FabULOUS, highlighting the versatility of the SmartEnzymes family.
Gstöttner et al., CE-MS analysis of highly homologous bispecifics.
The nature of the sheathless CE approach meant that the authors could collect and compare both the intact and subunit data using the same instrument setup. From that, marco- and microheterogeneity could be assessed. Differences between the two BsAb could be detected such as incomplete assemblies and free chains, along with PTMs such as glycation. Combining the observations, the authors conclude that the different engineering processes for each BsAbs results in varying heterogeneity. They also underscore the wide applicability of sheathless CE. Finally, they reveal the importance of SmartEnzymes and subunit analysis.

Easy IgG Isotype Fingerprinting with FabRICATOR® and Ion-Mobility MS
Researchers at Strasbourg University and Pierre-Fabre highlight the benefits of a middle-level approach to IgG isotype fingerprinting using native ion-mobility mass spectrometry.
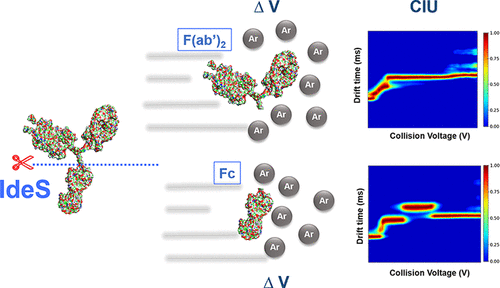
In this new article by Botzanowski et al., a middle-level approach was compared to more classical intact methods for distinguishing IgG isotypes using native ion mobility mass spectrometry. Ion mobility mass spectrometry and collision induced unfolding (CIU) at the intact level is hampered by minimal variations that can be observed. The authors used FabRICATOR to digest adalimumab, panitumumab, and natalizumab down to Fc/2 and F(ab’)2 domains. The Fc domain provided only limited isotype information due to sequence similarity. However, the stability profile of F(ab’)2 and its’ unfolding pattern measured by CIU uncovered very clear differences between the isotypes that could not be achieved with full length, intact mAb.
Eculizumab, a humanized IgG2/4 hybrid, which gives conflicting isotype patterns using classical approaches, was also tested. With the middle-level CIU approach, eculizumab could be easily distinguished from reference isotypes. In summary, the authors clearly show how easy, reliable and clear-cut classification of mAb isotypes can be achieved using a middle-level approach with FabRICATOR digestion.
The full article is available here:

Fully automated 3D LC-MS workflow for the characterization of antibody glycosylation
Genentech used FabRICATOR®-HPLC to develop a fast and automated 3D LC-MS workflow for the characterization of glycosylation on therapeutic antibodies.
The glycosylation profile is a critical quality attribute of biopharmaceuticals including antibodies. This heterogenous quality attribute can be tackled using a range of different analytical technologies including separation of released glycans , intact glycoprotein mass analysis, subunit mass analysis and many more. An antibody subunit approach using FabRICATOR (IdeS) combined with separation on hydrophilic interaction chromatography (HILIC) offers a promising strategy for characterization of IgG glycosylation. Typically, the FabRICATOR digestion is done manually, offline, prior to LC-MS. In a growing number of cases, like online monitoring of glycosylation, it is desirable to automate such a workflow.
Researchers at Genentech in collaboration with University of Geneva recently published an automated workflow using three dimensional separation coupled to mass spectrometry. To automate the workflow, they employed the FabRICATOR-HPLC column to deliver online antibody subunit generation. In their setup, this was followed by on column reduction and separation of the subunits using a reversed phase column, coupled to HILIC column to separate the Fc/2 glycoforms. In summary, the system incorporated three columns controlled by two valves: FabRICATOR-HPLC x Reduction/RPLC x HILIC/MS (Fig. 1 below).
Camperi et al. compared their automated workflow to the standard manual digestion by testing two of their candidate mAbs. The authors concluded that the 3D automated workflow, which took 95 minutes per sample, could deliver comparable data for all N-glycan variants for the Fc/2 subunit and therefore could be applied to the analysis of not only mAbs but also antibody drug conjugated (ADCs) and bispecific antibodies. Finally, the scientists emphasize that the key strength of the 3D approach was not just the automation aspect but also the fact that no tedious sample preparation was required, and that it is applicable to routine analysis tasks.

Fig 1. 3D-LC Workflow for automated middle-up analysis of mAbs (Camperi et al., 2020.)

SmartEnzymes at BGI, an interview with Aaron Bailey
We got the opportunity to talk to Aaron Bailey at BGI Americas. From their newly setup MS labs, they use SmartEnzymes to help provide analytical services to their customers globally. Read on to find out more.
Tell us about yourself and BGI?
Currently I work at BGI Americas as the Associate Director of Product Management for Mass Spectrometry Services. I am part of the scientific team based in San Jose, California, at the newly built Mass Spectrometry Center.
How has the MS team set up the industrial platform there at BGI in San Jose?
Our mass spec center is a service and R&D laboratory focused on providing LC-MS solutions for proteomic research and biologic drug characterization. Our current LC-MS service platform plays to our core strengths in automated sample prep, multi-mode HPLC separations, high resolution Orbitrap MS, and advanced bioinformatic analysis. We have built a versatile system which can cater to a wide scope of proteomics and biologic drug characterization needs. In San Jose our Q Exactive HF-X BioPharma can be coupled to either nano-flow or analytical flow UHPLC, which essentially affords us the ability to provide LC-MS services for multi-omic researchers and pharmaceutical research and development.
How does the MS team do sample preparation prior to MS analysis?
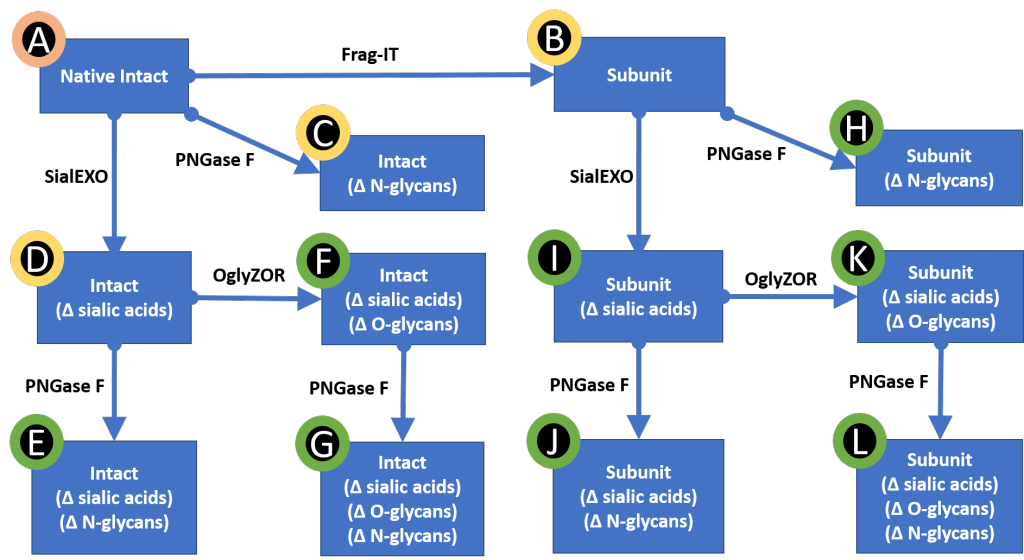
For emerging formats, including some Fc-fusion protein drugs which are already established in global markets, glycosylation can be both highly heterogeneous and often involve both N- and O-linkages and is often distributed on potentially many sites throughout the amino acid sequence. These projects require that glycosylations can be selectively removed in order to validate the identity of each identified PTM. We have had success using several deglycosidases including SmartEnzymes from Genovis. In a sort of combinatorial sample prep matrix these enzymes help objectively determine for each therapeutic protein how highly complex glycoform patterns obseved at the (completely unprocessed) intact mass level are specifically derived from the contributions of possibly several types of glycosylation events (i.e., N-linked vs. O-linked vs. sialylated). Each may individually involve heterogeneous distibution of the specific sugar types found at any particular amino acid site. While protein isoform distributions must be measured at the “intact” or “subunit” level, for example using FabRICATOR, site specific measurements must be performed using peptide mapping approaches. Most importantly, the two types of datasets must essentially agree on the findings, and the sample prep strategy described here is a powerful way to navigate this challenge.
What are the advantages of using SmartEnzymes like FragIT, OglyZOR and SialEXO?
We found that the activity and purity of these SmartEnzymes was a good fit for our native LC-MS platform in the sense that we are able to efficiently deglycosylate our therapeutic proteins using very little amount enzyme which remains near or below the actual level of detection in our experiments. This means that in normal usage we are not seeing these enzymes (FragIT, OglyZOR and SialEXO) in our datasets, which is particularly important for our native LC-MS intact mass analyses. This is a critical point for us as it means that we can keep our analyses streamlined by avoiding the need to address any artifact data which are not specifically related to any therapeutic protein isoforms.
The San Jose Mass Spectrometry Center uniquely allows BGI to offer state-of-the-art genomic and proteomic analytical services globally. In the coming year we plan to grow our team and add new LC and MS instrumentation to further increase our ability to support this market. We expect to see exciting new developments in integrating NGS and LC-MS datasets. We also plan to introduce additional mass spectrometry services which will allow us to provide even deeper support in our main focus areas of proteomics and biologics characterization.
Download Aaron’s ASMS poster here.
For more information on FragIT, OglyZOR and SialEXO please visit the following pages:
- FragIT, OglyZOR and SialEXO Product Pages
- Generation of Antibody Fragments & Analyis of O-glycans


FabRICATOR® in capillary zone electrophoresis-native mass spectrometry
The use of capillary electrophoresis for the analysis of therapeutic antibodies and other biopharmaceuticals is growing in popularity. A new article by researchers from CNRS in France showed how capillary zone electrophoresis-native mass spectrometry could be used for the quality control of intact therapeutic monoclonal antibodies. The authors show for the first time the use of a triple layer coated (PB-DS-PB) capillary with mAbs, which helps prevent mAbs adsorption. The intact therapeutic mAb, Infliximab, was analyzed under non-denaturing conditions to retain conformational heterogeneity and avoid denaturation. A middle up approach using FabRICATOR digestion was used to characterize dimer association detected in the stressed mAbs preparation. Digestion below the hinge region of the mAb produced F(ab’)2 and Fc/2 fragments which was subsequently analysed by capillary zone electrophoresis-native mass spectrometry. Using this approach the authors were able to see nature of the dimer association while maintaining the non-covalent interactions of the Fc/2 fragments.

For more information on FabRICATOR please visit the following pages:
- FabRICATOR Product Page
- Generation of Antibody Fragments
FabRICATOR® driven middle-down glycan analysis using NMR

Analysis of the glycosylation of therapeutic antibodies and other biopharmaceuticals is typically done by LC or LC/MS-based methods. However, each analytical technique has its strengths and weaknesses which makes the development of orthogonal methodologies crucial for in-depth characterization. In this study, researchers from the FDA present a middle-down NMR approach for studying glycosylation of therapeutic antibodies. Analogous to middle-down MS methods, the antibodies were digested using FabRICATOR to yield Fc/2 fragments. After chemical denaturation, these exhibited high enough solubility and sufficiently fast molecular dynamics to allow for glycan analysis by 2D-NMR without the need for isotope labeling or glycan release. Using this method, the authors were able to determine the chemical structure, glycosidic linkage position and anomeric configuration of each monosaccharide unit of the major Fc N-glycan structures and were able to quantify important quality attributes such as galactosylation and fucosylation.

For more information on FabRICATOR please visit the following pages:
- FabRICATOR Product Page
- Generation of Antibody Fragments

SmartEnzymes™ assist MALDI in-source decay FT-ICR Mass Spectrometry analysis
The Consortium for Top-down Proteomics is currently conducting a large inter lab study. They are developing methods for the analysis of intact mAbs and antibody subunits generated by digestion with either FabRICATOR or GingisKHAN. In this paper, van der Burgt et al. present a novel method for the analysis of mAbs based on MALDI in-source decay fragmentation coupled with high resolution FT-ICR mass spectrometry. The standard method for antibody sequence confirmation by MS is based on fragmentation using electron transfer dissociation (ETD). The MALDI-ISD based method yielded complementary fragments to those observed in ETD experiments, translating to increased sequence coverage. Using digestion with either FabRICATOR or GingisKHAN, a higher total sequence coverage could be achieved than for the intact mAbs. FabRICATOR digestion also allowed for direct analysis of Fc glycosylation by MALDI-FT-ICR without the need for LC separation.
Meet the Scientist
We got the opportunity to interview the last author of the paper, Dr Simone Nicolardi at Leiden University Medical Center.
Tell us about yourself?

I am a senior researcher at the Center for Proteomics and Metabolomics at Leiden University Medical Center (https://www.lumc.nl/org/proteomics-metabolomics/). My work is focused on the development of ultrahigh resolution mass spectrometry-based methods for the analysis of biomolecules such as glycans, peptides, and proteins. This includes the structural characterization of monoclonal antibodies for determination of primary amino acid sequence and post-translational modifications.
What is new with the MALDI in-source decay fragmentation you have published?
Our newly developed method combines the advantage of ultrahigh resolution mass measurements, obtained using high-end instrumentation and novel spectra processing software, with the efficient fragmentation provided by MALDI-in-source decay using 1,5-diaminonaphthalene as a reducing MALDI matrix. The main advantages are complementary sequence information compared to other mass spectrometric fragmentation techniques and the use of minimal sample preparation procedure that does not require separation techniques such as liquid chromatography.
What are the advantages of using FabRICATOR or GingisKHAN?
In MALDI-ISD experiments singly charged ions are generated from the protein backbone. Thus, a wide m/z-range is needed for the detection of all fragments generated from large compounds, such as monoclonal antibodies. Even in high-end MS instrumentation, this m/z-range is limited and sequence information is obtained from N- and C-terminal protein portions only. The use of FabRICATOR or GingisKHAN allows extending sequence information of heavy chains to more internal protein regions. In addition, after digestion mAb Fc portion can be analyzed directly by MALDI-MS allowing the detection of most abundant Fc glycoforms.
How do you view sample preparation prior to MS analysis of mAbs?
The structure characterization of mAbs is generally performed using a multi-level approach based on different analytical methods. Many of these methods are based on liquid chromatography (LC) electrospray ionization (ESI) MS. LC is used to separate mAb subunits and to remove ESI-non-compatible compounds such as salts. Our method is based on MALDI which is known to be more tolerant to salts in the sample. Thus, MALDI-ISD MS avoids laborious sample preparation steps that can lead to artificial modification of mAbs. Also, it comes with short analysis times in contrast to other chromatographic techniques.
What are you working on now?
Our attention is now on bispecific mAbs. We apply our MALDI-based method for the simultaneous analysis of all six different polypeptide chains generated after treatment with FabRICATOR and chemical reduction of disulfide bonds. Our aim is to develop a fast method for the monitoring of chemically induced mAb modifications.

For more information on FabRICATOR or GingisKHAN please visit the following pages:
- FabRICATOR and GingisKHAN Product Page
- Generation of Antibody Fragments