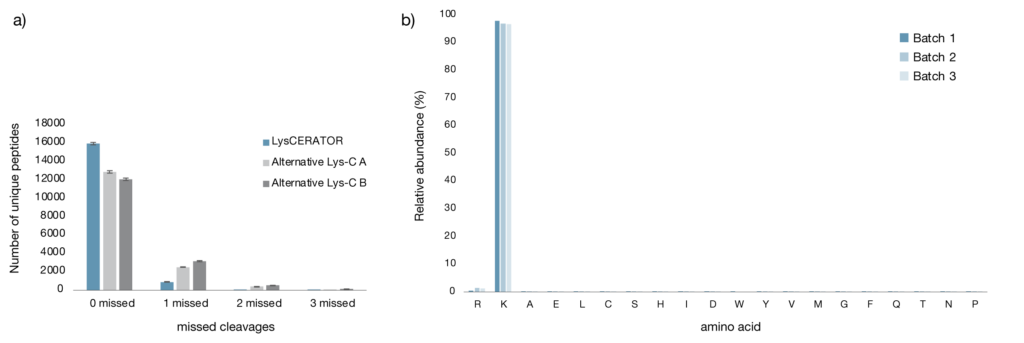Lysine-specific Protein Digestion
Page Specific Links
- Superior Digestion Efficiency of LysCERATOR on HeLa Cell Lysates
- Improved Digestion Efficiency of LysCERATOR on hIgG1
- LysCERATOR Enhances Digestion Efficiency at K-P Sites
- LysCERATOR Retains High Activity Under Strong Denaturing Conditions
- LysCERATOR is Highly Resistant to Autoproteolysis
Site Links
Superior Digestion Efficiency and Cleavage at Lysine-Proline Sites
LysCERATOR (Lys-C) is a lysyl endopeptidase that specifically digests peptide bonds C-terminally to lysine residues, including at lysine-proline linkages.
Lys-C is a widely used protease in proteomics and biopharmaceutical workflows, valued for its strict lysine specificity, high catalytic efficiency, and minimal autoproteolysis. These properties enable controlled and reproducible protein digestion, even under strongly denaturing conditions. Building on these defining characteristics, LysCERATOR offers enhanced performance, delivering superior digestion efficiency, particularly at challenging lysine–proline linkages, while maintaining robustness and precision in demanding workflows. These qualities make LysCERATOR especially well-suited for applications requiring lysine-specific digestion that demand precision, consistency, and stringent enzyme reliability.
![]()
Superior Digestion Efficiency of LysCERATOR™ on HeLa Cell Lysates
In proteomics applications, the choice of proteolytic enzyme is critical for achieving confident protein identification and comprehensive proteome coverage. High enzymatic efficiency is essential to digest complex protein mixtures into peptides suitable for LC–MS analysis. This increases peptide yield, improves sequence coverage, and enhances the likelihood of identifying low-abundance proteins. In this context, a high number of unique peptides - particularly peptides generated without any missed cleavages - is a key indicator of enzyme performance, as it directly contributes to increased confidence in consistent protein identification and quantification.
To evaluate digestion performance in a proteomics setting, human HeLa cell lysates were digested using LysCERATOR and two alternative Lys-C enzymes (Alternative Lys-C A and Alternative Lys-C B, Fig. 1a) and the numbers of unique peptides were compared. Following digestion, peptides were cleaned up using protein aggregation capture (PAC) on magnetic beads and analyzed by LC–MS. LysCERATOR generated both the highest number of unique peptides and the highest proportion of peptides containing zero missed cleavages of the three Lys-C enzymes tested. The higher proportion of fully cleaved peptides using LysCERATOR indicates superior digestion efficiency on complex biological samples, resulting in a more informative and interpretable peptide pool compared to the alternative enzymes.
Enzyme specificity is another critical parameter in proteomics workflows. A high level of amino acid specificity ensures predictable cleavage patterns and reduces the number of nonspecific peptides. This, in turn, minimizes spectral complexity and decreases ambiguity during peptide-to-spectrum matching. Three different batches of LysCERATOR were evaluated, and the lysine specificity was consistently high, at approximately 97–98% (Fig. 1b). The observed batch-to-batch reproducibility highlights the robustness and reliability of LysCERATOR for proteomics workflows.
LysCERATOR combines very high lysine specificity with superior digestion efficiency, making it well suited for complex proteomics workflows. These features result in cleaner spectra, greater confidence in protein identification, and more robust data analysis – particularly for highly complex samples with inherently large numbers of potential peptide assignments.
Figure 1. a) Comparison of Lys-C digestion of HeLa cells. HeLa cell lysate was digested in triplicate, with LysCERATOR and two alternative Lys-C enzymes, at an enzyme to substrate ratio of 1:100 in 50 mM HEPES buffer (pH 8.5) overnight at 37°C. Peptide clean-up was performed with PAC on magnetic beads and data were acquired by LC-MS. Graphs show the number of unique peptides identified grouped by the number of missed cleavages. Error bars show the standard deviation. b) Lysine specificity of LysCERATOR. Three different batches of LysCERATOR were used to digest HeLa cell lysate at an enzyme to substrate ratio of 1:100 in 50 mM HEPES buffer (pH 8.5) overnight at 37°C. Peptide clean-up was performed with PAC on magnetic beads and data were acquired by LC-MS. The amino acid specificity of the different batches of LysCERATOR was determined based on the frequency of the C-terminal amino acid from the identified peptides.
![]()
Improved Digestion Efficiency of LysCERATOR™ on hIgG1
High digestion efficiency is critical for reliable peptide mapping of antibody candidates in drug development. Peptide mapping is routinely used to confirm primary sequence, identify post-translational modifications (PTMs), and monitor product quality throughout development and manufacturing. Incomplete or inconsistent proteolytic digestion leads to missed cleavages, which increase sample complexity, reduce sequence coverage, and complicate data interpretation. This can obscure critical quality attributes (CQAs), reduce confidence in PTM identification and localization, and negatively impact comparability studies and regulatory submissions.
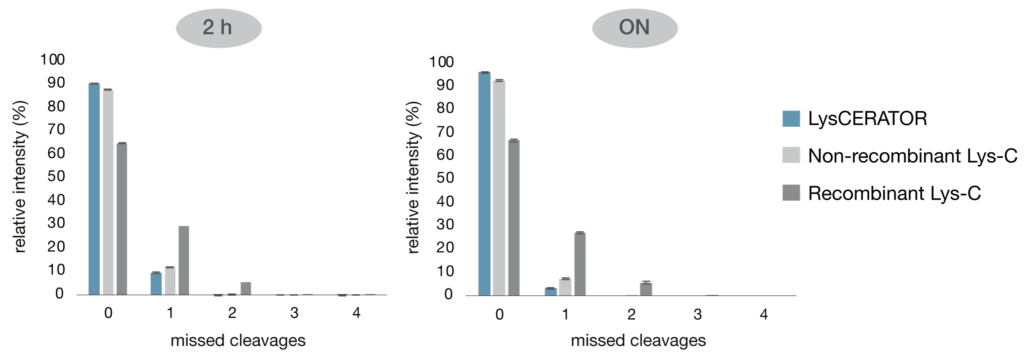
To evaluate the performance of LysCERATOR for peptide mapping of antibody candidates, the human IgG1 trastuzumab was digested using LysCERATOR and two alternative, commercially available Lys-C enzymes - one non-recombinant and one recombinant (Fig. 2). Following a 2-hour digestion, LysCERATOR achieved superior cleavage efficiency, with 91% of total peptide intensity corresponding to peptides with zero missed cleavages, compared to 88% for the non-recombinant Lys-C and 65% for the recombinant Lys-C. Following overnight digestion, the proportion of peptides with zero missed cleavages increased to 97% for LysCERATOR and 93% for the non-recombinant Lys-C, while remaining substantially lower at 67% for the recombinant alternative Lys-C.
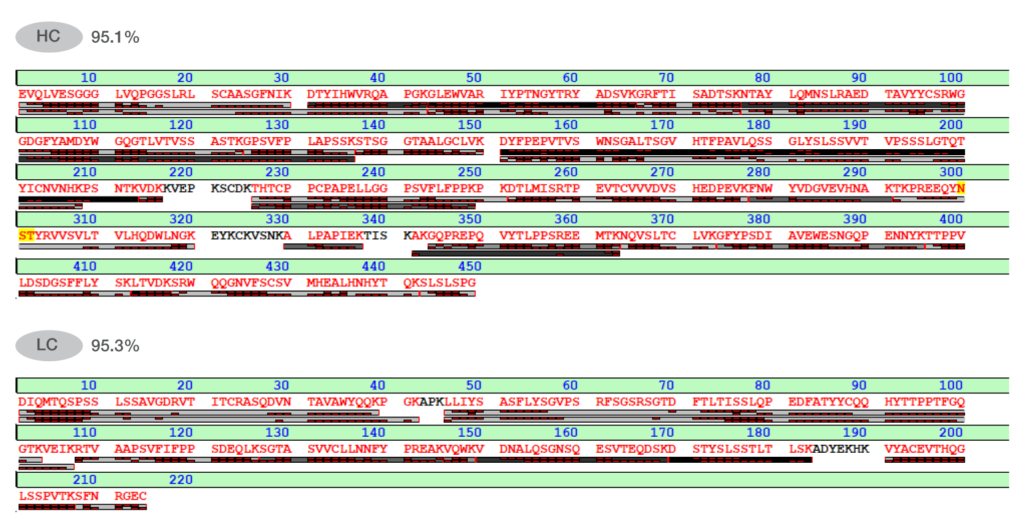
These results highlight the superior digestion efficiency of LysCERATOR for peptide mapping workflows compared to other commercially available products. Efficient digestion, with minimal missed cleavages, even at relatively short digestion times, enables faster turnaround without compromising data quality. This is particularly valuable in high-throughput analytical environments and early-stage development, where speed and reproducibility are essential. To further demonstrate the utility of LysCERATOR for peptide mapping, trastuzumab sequence coverage was assessed (Fig. 3). After 2 hours, sequence coverage reached 95.1% for the heavy chain and 95.3% for the light chain. High sequence coverage increases confidence in the detection and quantification of post-translational modifications when using LysCERATOR, which is critical during the comprehensive characterization of antibody therapeutics.
Figure 2. Comparison of Lys-C digestion of human IgG1. Trastuzumab was denatured, reduced and alkylated before digestion with LysCERATOR, a commercially available non-recombinant Lys-C and a commercially available recombinant Lys-C. Triplicate digestions with each enzyme were performed in 2 M of urea at an enzyme to substrate ratio of 1:50 in 0.1 M Tris buffer (pH 8.0) for 2 hours or overnight at 37°C. Data were acquired by LC-MS and processed using BioPharmaCompass. Bars of the graphs represent the mean relative peptide intensity for each number of missed cleavages across the triplicate digestions, with error bars indicating the standard deviation.
![]()
LysCERATOR™ Enhances Digestion Efficiency at K-P Sites
Cleavage at lysine residues followed by proline (K–P sites) is a well-established challenge for Lys-C digestion, often leading to missed cleavages due to proline-induced conformational constraints. To assess how effectively LysCERATOR overcomes this limitation, its performance at K–P sites was evaluated in detail and compared with two alternative, commercially available Lys-C enzymes - one non-recombinant and one recombinant.
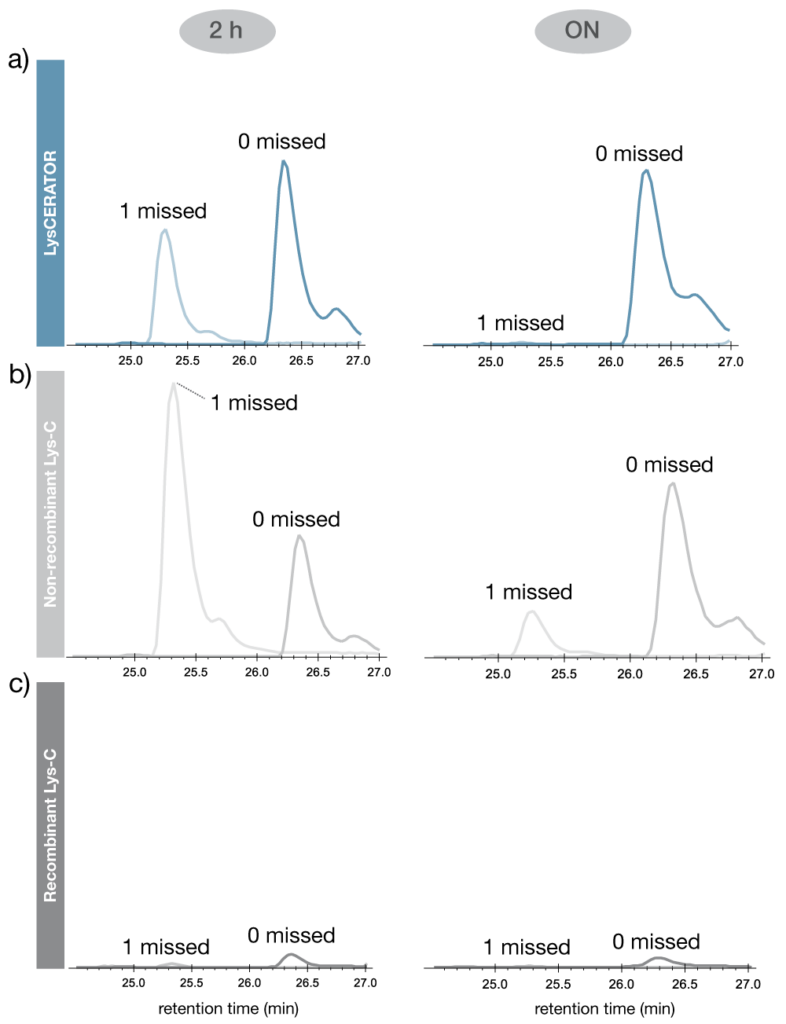
Digestion efficiency was assessed using trastuzumab. Figure 4 shows the relative intensities of two peptides derived from the light chain N-terminus: LC aa 1–39, representing the expected cleavage, and LC aa 1–42, which contains a missed cleavage at the K39/P40 site. After 2 hours of digestion with LysCERATOR, the fully cleaved peptide (LC aa 1–39) was the more abundant species, while digestion with the non-recombinant alternative Lys-C predominantly produced the miscleaved peptide (LC aa 1–42).
Following overnight digestion with LysCERATOR, there was a near-complete digestion at the K-P site while, in contrast, a substantial amount of the miscleaved peptide remained after digestion with the non-recombinant alternative Lys-C. Digestion with the recombinant alternative Lys-C resulted in low overall signal intensity for both peptides, with the most prominent species being a peptide containing two missed cleavages (LC aa 1–45 – not shown).
Collectively, these results demonstrate that LysCERATOR exhibits superior digestion efficiency at challenging K–P sites compared with alternative Lys-C enzymes, resulting in fewer missed cleavages and more complete proteolytic processing.
Figure 4. Comparison of Lys-C digestion efficiency at K-P sites. Intensities of two trastuzumab light chain peptides 1) aa 1-39, representing the expected cleavage product, and 2) aa 1-42, containing a missed cleavage at the K39/P40 site with a) LysCERATOR , b) Non-recombinant alternative Lys-C or c) Recombinant alternative Lys-C. Trastuzumab was denatured, reduced and alkylated before digestion in 2 M of urea at an enzyme to substrate ratio of 1:50 in 0.1 M Tris buffer (pH 8.0) for 2 hours or overnight at 37°C. Data were acquired by LC-MS and processed by searching the Mascot probability based search engine against the trastuzumab sequence followed by quantitative analysis in Skyline (MacCross lab).
![]()
LysCERATOR™ Retains High Activity Under Strong Denaturing Conditions
Efficient proteolytic digestion generally requires denaturation of the target protein to fully expose potential cleavage sites. However, some proteins are highly structured, or aggregation-prone, and, therefore, may require high concentrations of denaturing agents to render the protein backbone accessible for enzymatic digestion. To evaluate the robustness of LysCERATOR under such challenging conditions, its digestion performance was assessed in the presence of increasing concentrations of commonly used denaturants.
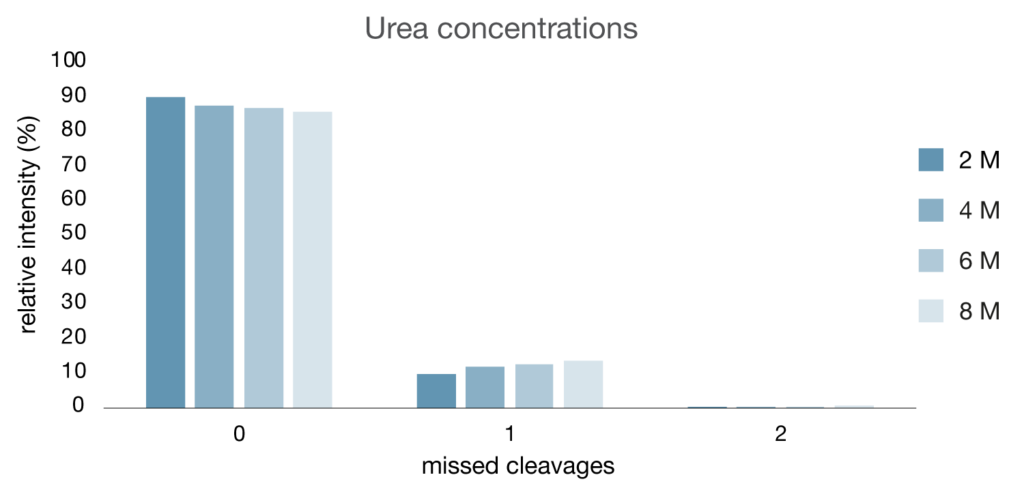
Trastuzumab was digested with LysCERATOR across a range of urea concentrations (Fig. 5), and digestion efficiency was largely preserved between 2–8 M urea. After 2 hours digestion at 37°C, the relative intensity of peptides with zero missed cleavages remained high, decreasing only slightly from 90% at 2 M urea to 86% at 8 M urea, with little impact on overall peptide intensity. This indicates that even a high concentration of urea has minimal impact on LysCERATOR performance.
The performance of LysCERATOR was also evaluated in the presence of guanidine hydrochloride (Gd-HCl). Efficient digestion was maintained between 1–2 M Gd-HCl, with the relative intensity of peptides with zero missed cleavages remaining around 90% after 2 hours at 37°C (data not shown). At higher denaturant strength (4 M Gd-HCl), the proportion of peptides with zero missed cleavages decreased to 68%, reflecting reduced digestion efficiency consistent with partial inhibition of enzymatic activity under these conditions.
Together, these results demonstrate that LysCERATOR exhibits a high tolerance to a strong denaturing environment when using urea, and moderate concentrations of Gd-HCl. This robustness highlights its suitability for proteomic analysis of proteins that are difficult to denature, where strongly denaturing concentrations are required to achieve comprehensive and reproducible digestion.
Figure 5. Impact of urea concentration on LysCERATOR performance. Trastuzumab was digested with LysCERATOR across a range of 2-8 M of urea. The antibody was denatured, reduced and alkylated prior to digestion at an enzyme to substrate ratio of 1:100 in 0.1 M Tris buffer (pH 8.0) for 2 hours at 37°C. Data were acquired by LC-MS and processed using BioPharmaCompass (Bruker).
![]()
LysCERATOR™ is Highly Resistant to Autoproteolysis
A low level of autoproteolysis is an important quality attribute of proteolytic enzymes used for peptide mapping of antibody candidates in drug development. Autoproteolysis - self-digestion of the enzyme - can negatively affect analytical performance in two important ways. Firstly, it may reduce the effective enzyme activity during longer digestion times, leading to incomplete digestion and increased variability. Secondly, peptides generated from enzyme self-cleavage can co-elute with analyte-derived peptides and introduce contaminating signals during LC–MS analyses, complicating peak assignment and increasing the risk of misidentification. This added ambiguity is particularly problematic in peptide mapping workflows, where confident sequence confirmation and precise localization of post-translational modifications are essential for defining critical quality attributes (CQAs).
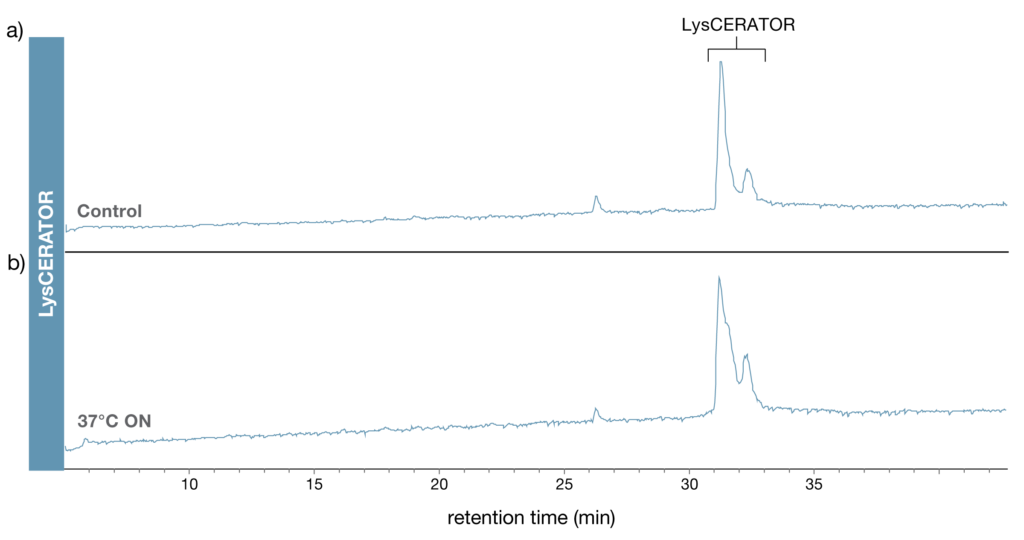
To investigate potential autoproteolytic tendencies of LysCERATOR, the enzyme was incubated overnight in reaction buffer (0.1 M Tris-HCl, pH 8.0) at 37°C and analyzed by RP-LC–MS (Fig. 6). A freshly prepared enzyme sample was analyzed in parallel as a control. No generation of LysCERATOR peptides was observed following incubation of the enzyme alone, compared to the control, consistent with very low or undetectable levels of LysCERATOR autoproteolysis under these conditions.
The absence of detectable autoproteolysis demonstrates the effectiveness of LysCERATOR for peptide mapping applications, particularly for extended or overnight digestions commonly used in antibody characterization. By minimizing enzyme-derived background signals and maintaining consistent proteolytic activity, low levels of autoproteolysis contribute to cleaner mass spectra, more confident peptide identification, and improved robustness of peptide mapping methods throughout antibody drug development.
Figure 6. LysCERATOR resistance to autoproteolysis. Samples of LysCERATOR were evaluated for potential autoproteolysis by RP-LC-MS using a C18 column. The total ion chromatograms of a) a freshly reconstituted LysCERATOR sample and b) a LysCERATOR sample which had been incubated overnight at 37°C in 0.1 M Tris-HCl (pH 8) are shown and indicate a lack of detectable LysCERATOR autoproteolysis under these conditions.